Tutorial guide of
TMMGdb - Tumor Metastasis Mechanism-associated Gene Database
1. Design principles
At present, all or the majority of the published databases classified cancer-associated genes based on cancer types. There are even fewer databases focused on collecting tumor metastasis mechanism-associated genes based on hallmarks of cancer/metastasis. In spite of intensive research, few published works address the problem of setting up a web-based resource on characterizing metastasis genes under the metastasis mechanism framework. Since tumor metastasis is a dynamic process and involving many cellular and molecular processes, those published databases classified metastasis genes based on either cancer types or hallmarks of cancer cannot provide information on the sequential relations as well as cellular and molecular mechanisms among different metastasis stages. In other words, there is no way such databases can provide a clear picture of the onset and progression of the tumor metastasis process. Therefore, TMMGdb provides a systematic approach for the biomedical community to gain an in-depth understanding of the dynamic process, molecular definitions and cellular processes involved in tumor metastasis. Compared with the state-of-art research work, TMMGdb is a novel, systematic and comprehensive resource.
2. Collection of tumor metastasis mechanism keywords
We utilized three methods to select tumor metastasis mechanism-associated “Level 1” and “Level 2” keywords. The three methods are: (a) keywords employed by literature using to build mathematical models of the cancer metastasis processes,(b) research/review papers addressing the metastasis process, and (c) hallmark gene set names (hallmarks of metastasis terms), extracted from The Molecular Signatures Database (MSigDB) (Liberzon et al., 2015). The “Level 1” keywords are relatively broad, whereas, “Level 2” keywords are more specific.
“Level 1” keywords – a total of four keywords: “primary detachment” OR "in situ metastasis", intravasation, motility OR migration OR extravasation and "distal metastasis" OR colonization OR proliferation.
The “Level 2” keywords comprises of 44 keywords, obtained from the MSigDB database, literature review and advice from two oncologists. The first 10 keywords (in alphabet order) are listed as follows:
“Level 2” keywords (the first 10 keywords): androgen, angiogenesis, apoptosis, autophagy, axonogenesis, cell adhesion, cell migration, chemokines, cholesterol, “chromosomal instability or genomic instability etc.
3. Input datasets
The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/) is one of the most comprehensive cancer research resources in the world. Genomic Data Commons (GDC) program research project is supported by TCGA National Cancer Institute (NCI). Information on the mechanism of tumor metastasis can be obtained from Broad Institute TCGA-GDAC Firebrowse. The Firebrowse database (http://firebrowse.org/) provides more than 80K samples from more than 11,000 cancer patients, involving 38 cancer diseases.
4. Search interfaces
Firstly, the TMMGdb database provides three interfaces; ‘Browse’, ‘Search’, ‘DEG search’ and ‘Download’, for users to retrieve information about TMMGs. TMMGdb contains a wealth of annotations, including: PubMed ID, genetics, cancer types, cancer cell lines, miRNAs, pathways and tissue expression, mutations, drug resistant, PPIN, PubMed and JCR journal ranking information.
Secondly, every TMMG can be referenced back to the PubMed IDs. The papers PubMed ID) displayed on the webpage are sorted according to the records of the JCR 2019 edition. It is convenient for users to select and read according to the ranking of the paper.
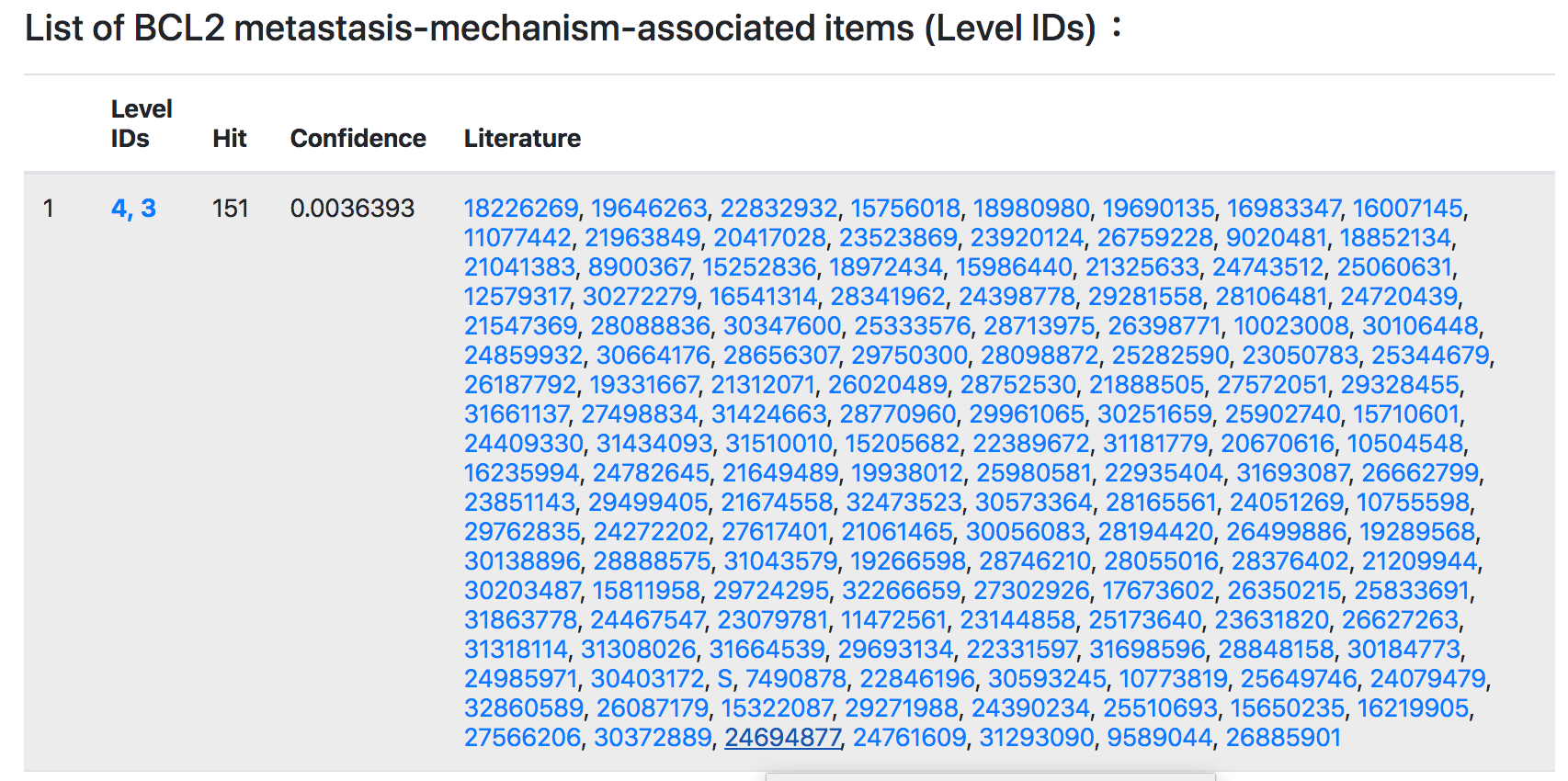
Regulatory pathway, miRNA regulators, tissue expression, cancer type and cancer cell lines information.
Thirdly, TMMGdb provides comprehensive annotations, including: pathway, miRNA regulators, tissue expression, cancer type annotation and cancer cell lines information.
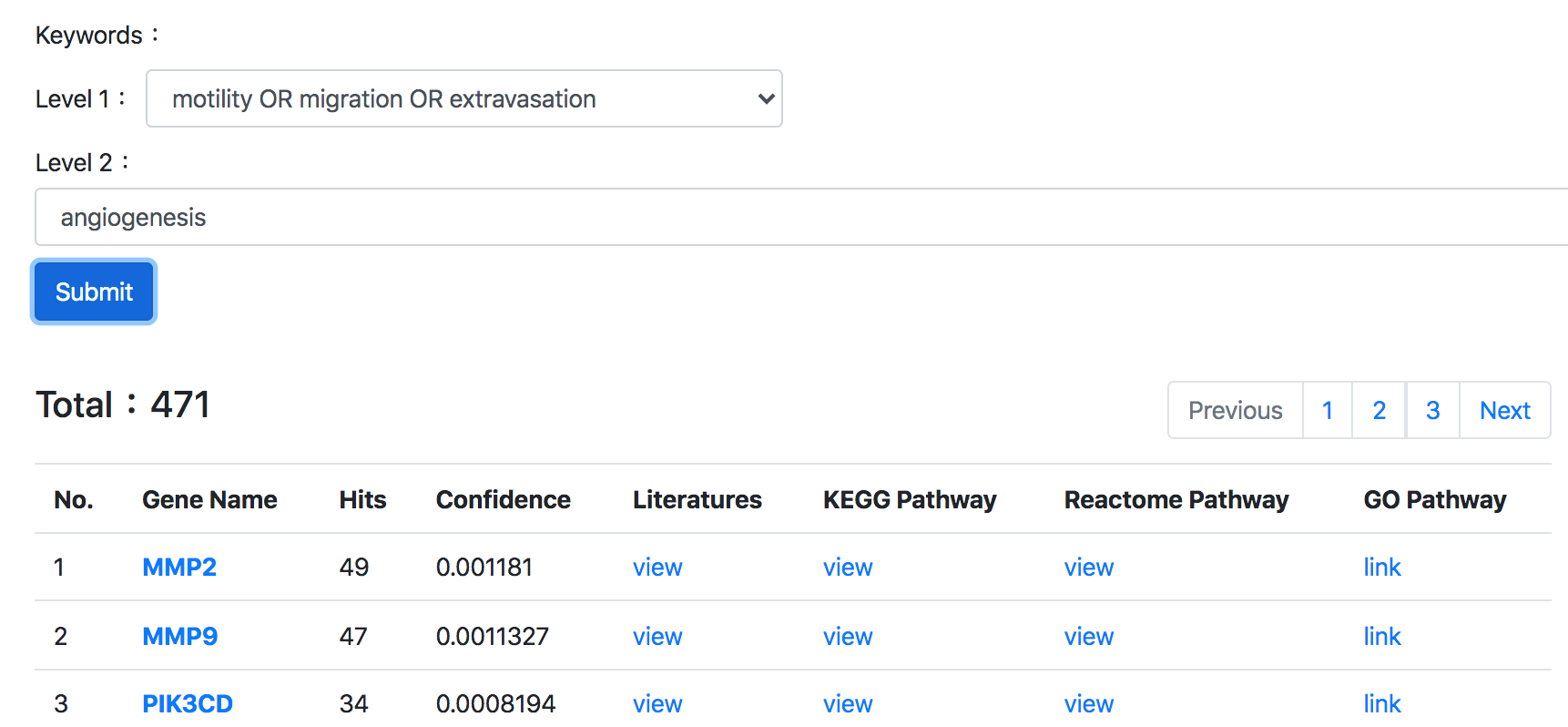


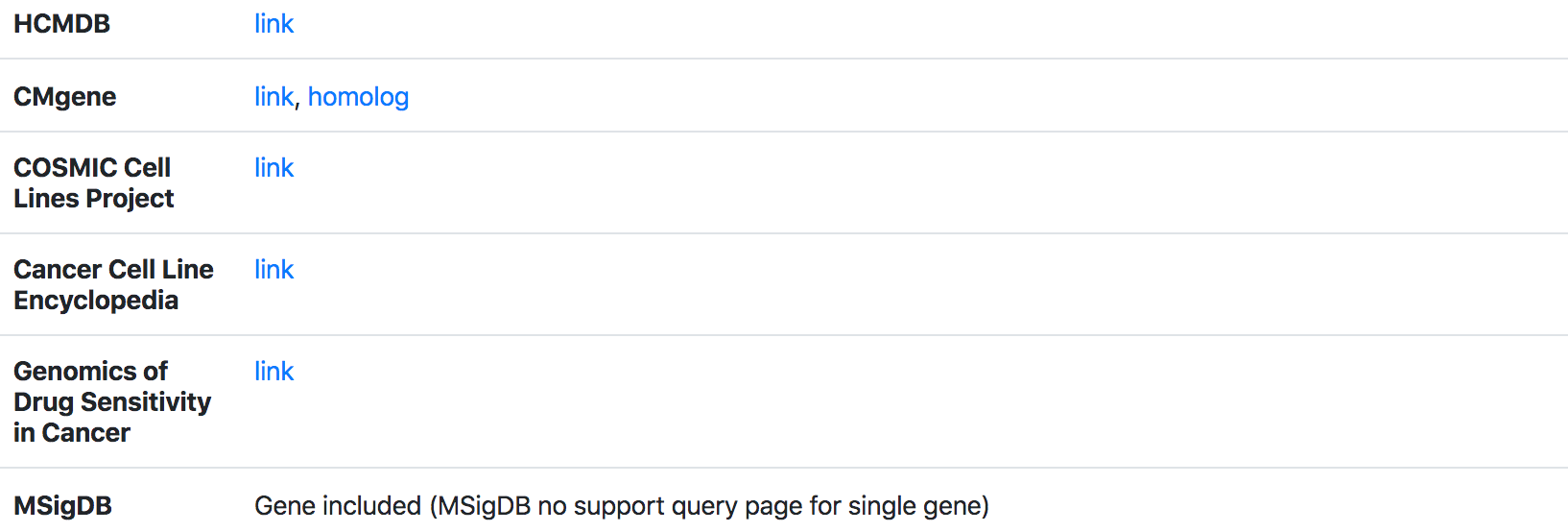
5. Genetic mutation and drug resistant annotations
Fourthly, TMMGdb provides mutation [including: driver gene, mutation type, translocation partner, hallmark, amino acid mutation, CDS mutation] and drug resistant annotation.
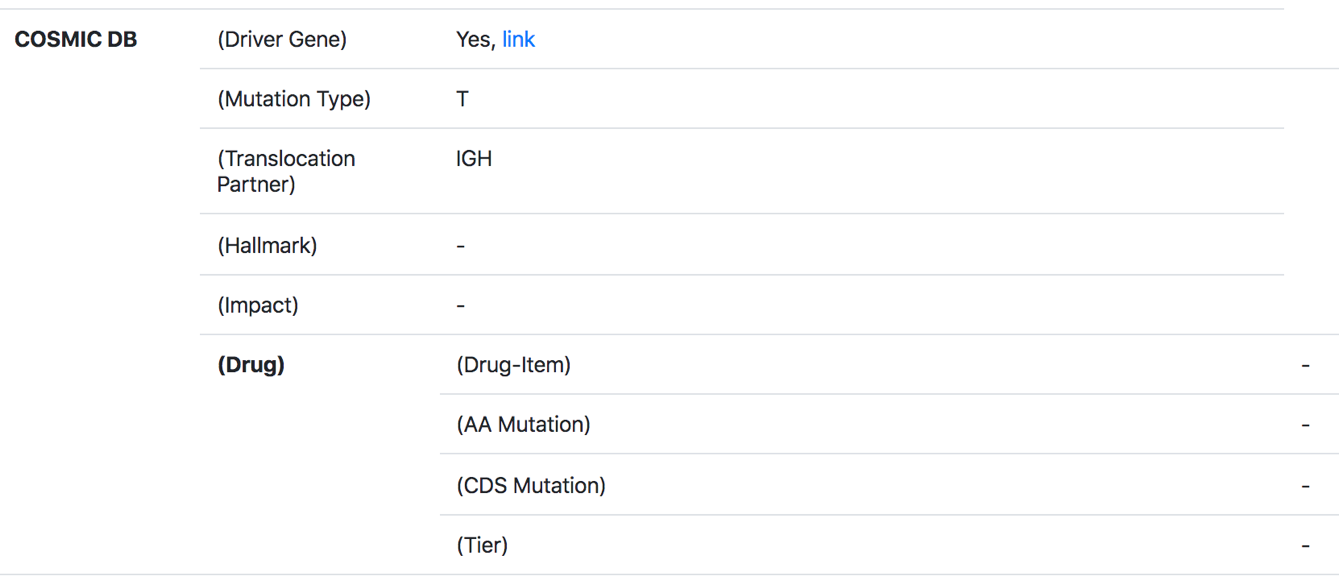
6. Transcription factor-mediated PPINs
Fifthly, TMMGdb provides transcription factor-mediated PPINs of all the TMMGs. In case the TMMG or its direct interacting partner is a TF, the activation or suppression regulatory information was provided by TRRUST.
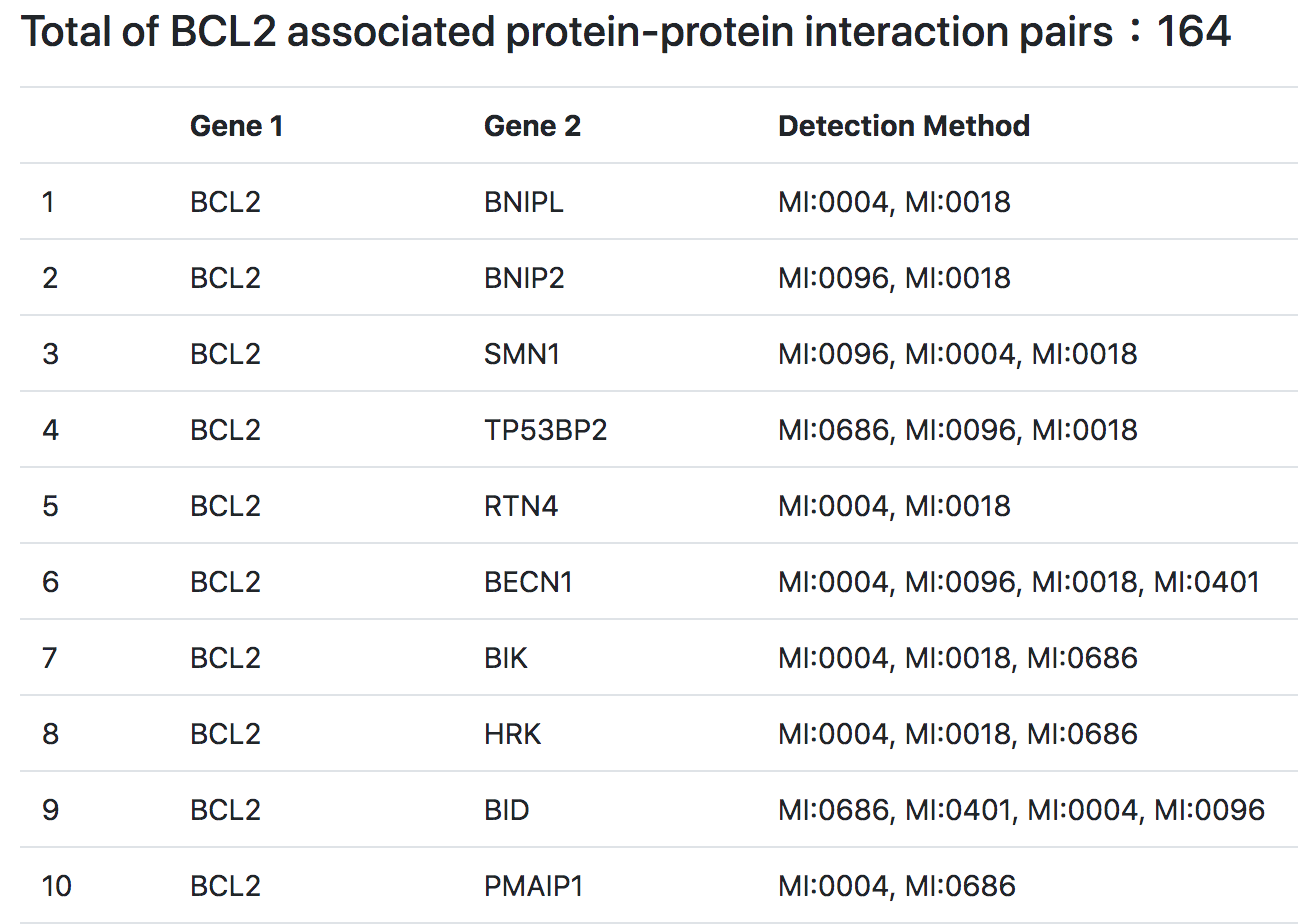
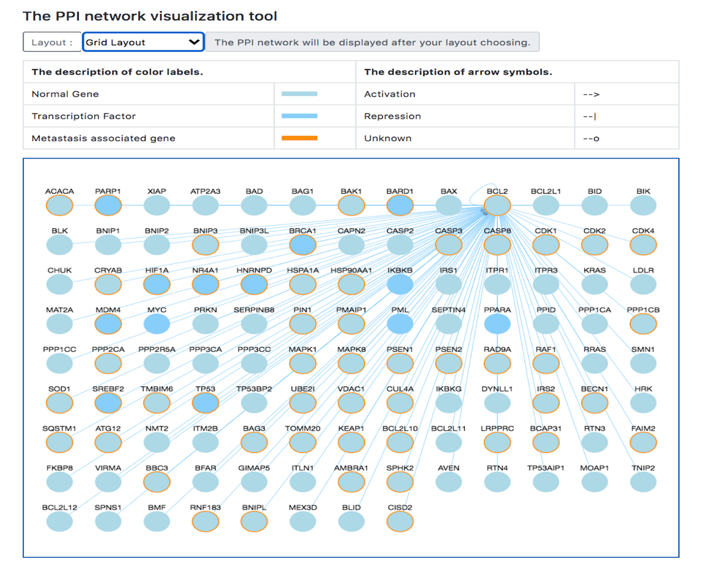
(A)
(B)
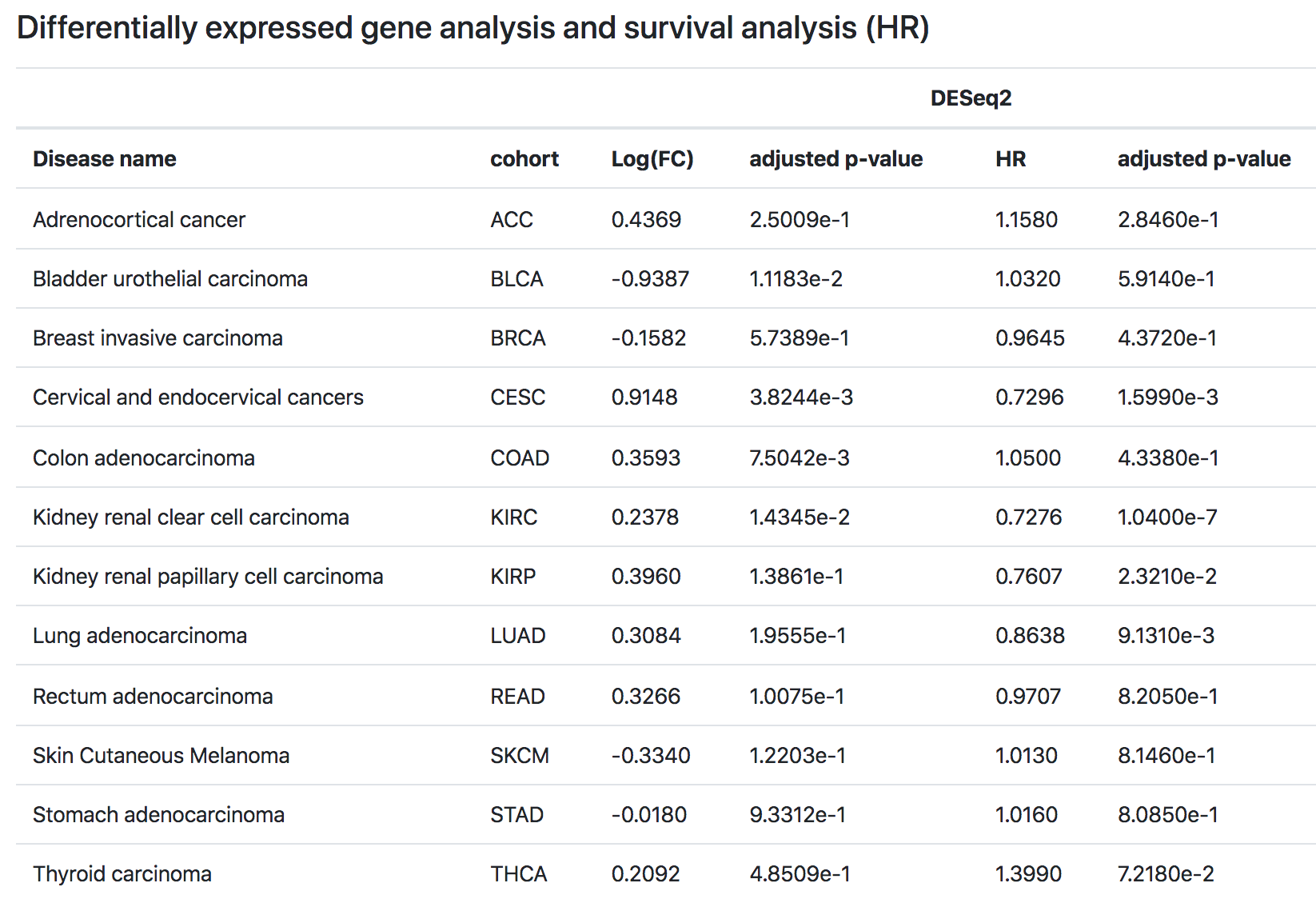
7. Differentially expressed gene analysis and survival analysis (HR)
Sixthly, TMMGdb provides differentially expressed genes (DEGs) results for 12 cancer cohorts. If the gene satisfies the Benjamini-Hochberg adjusted p-value less-than 0.05 and the absolute value of the logarithm of fold change (FC) greater than or equals to one (|Log2FC|>=1), we defined it as a DEG. DEGs associated with |Log2FC| >=1 and adj. p-value < 0.05 are potential biomarkers of tumor metastasis.
Copyright © 2021 Klng Lab., Asia University, Taiwan. All rights reserved.